13. Numba#
In addition to what’s in Anaconda, this lecture will need the following libraries:
!pip install quantecon
Please also make sure that you have the latest version of Anaconda, since old versions are a common source of errors.
Let’s start with some imports:
import numpy as np
import quantecon as qe
import matplotlib.pyplot as plt
13.1. Overview#
In an earlier lecture we discussed vectorization, which can improve execution speed by sending array processing operations in batch to efficient low-level code.
However, as discussed in that lecture, traditional vectorization schemes have weaknesses:
Highly memory-intensive for compound array operations
Ineffective or impossible for some algorithms
One way to circumvent these problems is by using Numba, a just in time (JIT) compiler for Python.
Numba compiles functions to native machine code instructions at runtime.
When it succeeds, the result is performance comparable to compiled C or Fortran.
In addition, Numba can do useful tricks such as multithreading.
This lecture introduces the core ideas.
Note
Some readers might be curious about the relationship between Numba and Julia, which contains its own JIT compiler. While the two compilers are similar in many ways, Numba is less ambitious, attempting only to compile a small subset of the Python language. Although this might sound like a deficiency, it is also a strength: the more restrictive nature of Numba makes it easy to use well and good at what it does.
13.2. Compiling Functions#
13.2.1. An Example#
Let’s consider a problem that’s difficult to vectorize (i.e., hand off to array processing operations).
The problem involves generating the trajectory via the quadratic map
In what follows we set \(\alpha = 4\).
13.2.1.1. Base Version#
Here’s the plot of a typical trajectory, starting from \(x_0 = 0.1\), with \(t\) on the x-axis
def qm(x0, n, α=4.0):
x = np.empty(n+1)
x[0] = x0
for t in range(n):
x[t+1] = α * x[t] * (1 - x[t])
return x
x = qm(0.1, 250)
fig, ax = plt.subplots()
ax.plot(x, 'b-', lw=2, alpha=0.8)
ax.set_xlabel('$t$', fontsize=12)
ax.set_ylabel('$x_{t}$', fontsize = 12)
plt.show()
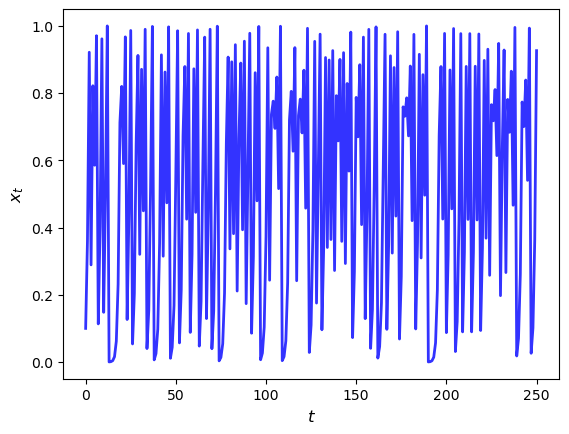
Let’s see how long this takes to run for large \(n\)
n = 10_000_000
with qe.Timer() as timer1:
# Time Python base version
x = qm(0.1, n)
4.6496 seconds elapsed
13.2.1.2. Acceleration via Numba#
To speed the function qm up using Numba, we first import the jit function
from numba import jit
Now we apply it to qm, producing a new function:
qm_numba = jit(qm)
The function qm_numba is a version of qm that is “targeted” for
JIT-compilation.
We will explain what this means momentarily.
Let’s time this new version:
with qe.Timer() as timer2:
# Time jitted version
x = qm_numba(0.1, n)
0.2020 seconds elapsed
This is a large speed gain.
In fact, the next time and all subsequent times it runs even faster as the function has been compiled and is in memory:
with qe.Timer() as timer3:
# Second run
x = qm_numba(0.1, n)
0.0716 seconds elapsed
Here’s the speed gain
timer1.elapsed / timer3.elapsed
64.94208656212433
This is a big boost for a small modification to our original code.
Let’s discuss how this works.
13.2.2. How and When it Works#
Numba attempts to generate fast machine code using the infrastructure provided by the LLVM Project.
It does this by inferring type information on the fly.
(See our earlier lecture on scientific computing for a discussion of types.)
The basic idea is this:
Python is very flexible and hence we could call the function qm with many types.
e.g.,
x0could be a NumPy array or a list,ncould be an integer or a float, etc.
This makes it very difficult to generate efficient machine code ahead of time (i.e., before runtime).
However, when we do actually call the function, say by running
qm(0.5, 10), the types ofx0,αandnare determined.Moreover, the types of other variables in
qmcan be inferred once the input types are known.So the strategy of Numba and other JIT compilers is to wait until the function is called, and then compile.
That is called “just-in-time” compilation.
Note that, if you make the call qm_numba(0.5, 10) and then follow it with qm_numba(0.9, 20), compilation only takes place on the first call.
This is because compiled code is cached and reused as required.
This is why, in the code above, the second run of qm_numba is faster.
Remark
In practice, rather than writing qm_numba = jit(qm), we typically use
decorator syntax and put @jit before the function definition. This is
equivalent to adding qm = jit(qm) after the definition.
13.4. Multithreaded Loops in Numba#
In addition to JIT compilation, Numba provides support for parallel computing on CPUs and GPUs.
The key tool for parallelization on CPUs in Numba is the prange function, which tells
Numba to execute loop iterations in parallel across available cores.
To illustrate, let’s look first at a simple, single-threaded (i.e., non-parallelized) piece of code.
The code simulates updating the wealth \(w_t\) of a household via the rule
Here
\(R\) is the gross rate of return on assets
\(s\) is the savings rate of the household and
\(y\) is labor income.
We model both \(R\) and \(y\) as independent draws from a lognormal distribution.
Here’s the code:
@jit
def update(w, r=0.1, s=0.3, v1=0.1, v2=1.0):
" Updates household wealth. "
# Draw shocks
R = np.exp(v1 * np.random.randn()) * (1 + r)
y = np.exp(v2 * np.random.randn())
# Update wealth
w = R * s * w + y
return w
Let’s have a look at how wealth evolves under this rule.
fig, ax = plt.subplots()
T = 100
w = np.empty(T)
w[0] = 5
for t in range(T-1):
w[t+1] = update(w[t])
ax.plot(w)
ax.set_xlabel('$t$', fontsize=12)
ax.set_ylabel('$w_{t}$', fontsize=12)
plt.show()
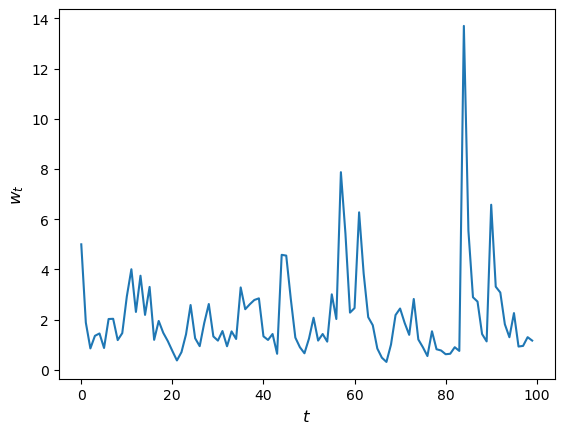
Now let’s suppose that we have a large population of households and we want to know what median wealth will be.
This is not easy to solve with pencil and paper, so we will use simulation instead:
Simulate a large number of households forward in time
Calculate median wealth
Here’s the code:
@jit
def compute_long_run_median(w0=1, T=1000, num_reps=50_000):
obs = np.empty(num_reps)
# For each household
for i in range(num_reps):
# Set the initial condition and run forward in time
w = w0
for t in range(T):
w = update(w)
# Record the final value
obs[i] = w
# Take the median of all final values
return np.median(obs)
Let’s see how fast this runs:
with qe.Timer():
# Warm up
compute_long_run_median()
6.6990 seconds elapsed
with qe.Timer():
# Second run
compute_long_run_median()
5.6500 seconds elapsed
To speed this up, we’re going to parallelize it via multithreading.
To do so, we add the parallel=True flag and change range to prange:
from numba import prange
@jit(parallel=True)
def compute_long_run_median_parallel(
w0=1, T=1000, num_reps=50_000
):
obs = np.empty(num_reps)
for i in prange(num_reps): # Parallelize over households
w = w0
for t in range(T):
w = update(w)
obs[i] = w
return np.median(obs)
Let’s look at the timing:
with qe.Timer():
# Warm up
compute_long_run_median_parallel()
1.0755 seconds elapsed
with qe.Timer():
# Second run
compute_long_run_median_parallel()
0.5861 seconds elapsed
The speed-up is significant.
Notice that we parallelize across households rather than over time – updates of an individual household across time periods are inherently sequential.
For GPU-based parallelization, see our lectures on JAX.
13.5. Exercises#
Exercise 13.1
Previously we considered how to approximate \(\pi\) by Monte Carlo.
Use the same idea here, but make the code efficient using Numba.
Compare speed with and without Numba when the sample size is large.
Solution
Here is one solution:
@jit
def calculate_pi(n=1_000_000):
count = 0
for i in range(n):
u, v = np.random.uniform(0, 1), np.random.uniform(0, 1)
d = np.sqrt((u - 0.5)**2 + (v - 0.5)**2)
if d < 0.5:
count += 1
area_estimate = count / n
return area_estimate * 4 # dividing by radius**2
Now let’s see how fast it runs:
with qe.Timer():
calculate_pi()
0.1860 seconds elapsed
with qe.Timer():
calculate_pi()
0.0322 seconds elapsed
If we switch off JIT compilation by removing @jit, the code takes around
150 times as long on our machine.
So we get a speed gain of 2 orders of magnitude by adding four characters.
Exercise 13.2
In the Introduction to Quantitative Economics with Python lecture series you can learn all about finite-state Markov chains.
For now, let’s just concentrate on simulating a very simple example of such a chain.
Suppose that the volatility of returns on an asset can be in one of two regimes — high or low.
The transition probabilities across states are as follows
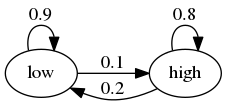
For example, let the period length be one day, and suppose the current state is high.
We see from the graph that the state tomorrow will be
high with probability 0.8
low with probability 0.2
Your task is to simulate a sequence of daily volatility states according to this rule.
Set the length of the sequence to n = 1_000_000 and start in the high state.
Implement a pure Python version and a Numba version, and compare speeds.
To test your code, evaluate the fraction of time that the chain spends in the low state.
If your code is correct, it should be about 2/3.
Hint
Represent the low state as 0 and the high state as 1.
If you want to store integers in a NumPy array and then apply JIT compilation, use
x = np.empty(n, dtype=np.int64).
Solution
We let
0 represent “low”
1 represent “high”
p, q = 0.1, 0.2 # Prob of leaving low and high state respectively
Here’s a pure Python version of the function
def compute_series(n):
x = np.empty(n, dtype=np.int64)
x[0] = 1 # Start in state 1
U = np.random.uniform(0, 1, size=n)
for t in range(1, n):
current_x = x[t-1]
if current_x == 0:
x[t] = U[t] < p
else:
x[t] = U[t] > q
return x
Let’s run this code and check that the fraction of time spent in the low state is about 0.666
n = 1_000_000
x = compute_series(n)
print(np.mean(x == 0)) # Fraction of time x is in state 0
0.66533
This is (approximately) the right output.
Now let’s time it:
with qe.Timer():
compute_series(n)
0.6052 seconds elapsed
Next let’s implement a Numba version, which is easy
compute_series_numba = jit(compute_series)
Let’s check we still get the right numbers
x = compute_series_numba(n)
print(np.mean(x == 0))
0.667017
Let’s see the time
with qe.Timer():
compute_series_numba(n)
0.0215 seconds elapsed
This is a nice speed improvement for one line of code!
Exercise 13.3
In an earlier exercise, we used Numba to accelerate an effort to compute the constant \(\pi\) by Monte Carlo.
Now try adding parallelization and see if you get further speed gains.
You should not expect huge gains here because, while there are many independent tasks (draw point and test if in circle), each one has low execution time.
Generally speaking, parallelization is less effective when the individual tasks to be parallelized are very small relative to total execution time.
This is due to overheads associated with spreading all of these small tasks across multiple CPUs.
Nevertheless, with suitable hardware, it is possible to get nontrivial speed gains in this exercise.
For the size of the Monte Carlo simulation, use something substantial, such as
n = 100_000_000.
Solution
Here is one solution:
@jit(parallel=True)
def calculate_pi(n=1_000_000):
count = 0
for i in prange(n):
u, v = np.random.uniform(0, 1), np.random.uniform(0, 1)
d = np.sqrt((u - 0.5)**2 + (v - 0.5)**2)
if d < 0.5:
count += 1
area_estimate = count / n
return area_estimate * 4 # dividing by radius**2
Now let’s see how fast it runs:
with qe.Timer():
calculate_pi()
0.4617 seconds elapsed
with qe.Timer():
calculate_pi()
0.0046 seconds elapsed
By switching parallelization on and off (selecting True or
False in the @jit annotation), we can test the speed gain that
multithreading provides on top of JIT compilation.
On our workstation, we find that parallelization increases execution speed by a factor of 2 or 3.
(If you are executing locally, you will get different numbers, depending mainly on the number of CPUs on your machine.)
Exercise 13.4
In our lecture on SciPy, we discussed pricing a call option in a setting where the underlying stock price had a simple and well-known distribution.
Here we discuss a more realistic setting.
We recall that the price of the option obeys
where
\(\beta\) is a discount factor,
\(n\) is the expiry date,
\(K\) is the strike price and
\(\{S_t\}\) is the price of the underlying asset at each time \(t\).
Suppose that n, β, K = 20, 0.99, 100.
Assume that the stock price obeys
where
Here \(\{\xi_t\}\) and \(\{\eta_t\}\) are IID and standard normal.
(This is a stochastic volatility model, where the volatility \(\sigma_t\) varies over time.)
Use the defaults μ, ρ, ν, S0, h0 = 0.0001, 0.1, 0.001, 10, 0.
(Here S0 is \(S_0\) and h0 is \(h_0\).)
By generating \(M\) paths \(s_0, \ldots, s_n\), compute the Monte Carlo estimate
of the price, applying Numba and parallelization.
Solution
With \(s_t := \ln S_t\), the price dynamics become
Using this fact, the solution can be written as follows.
M = 10_000_000
n, β, K = 20, 0.99, 100
μ, ρ, ν, S0, h0 = 0.0001, 0.1, 0.001, 10, 0
@jit(parallel=True)
def compute_call_price_parallel(β=β,
μ=μ,
S0=S0,
h0=h0,
K=K,
n=n,
ρ=ρ,
ν=ν,
M=M):
current_sum = 0.0
# For each sample path
for m in prange(M):
s = np.log(S0)
h = h0
# Simulate forward in time
for t in range(n):
s = s + μ + np.exp(h) * np.random.randn()
h = ρ * h + ν * np.random.randn()
# And add the value max{S_n - K, 0} to current_sum
current_sum += max(np.exp(s) - K, 0)
return β**n * current_sum / M
Try swapping between parallel=True and parallel=False and noting the run time.
If you are on a machine with many CPUs, the difference should be significant.